3 Simulation example
3.1 Data-generating process
The data-generating process is described via the directed acyclic graph (DAG) in Fig. 3.1. In this DAG, window shading \((\text{W})\) influences perceived warmth \((\text{P})\) both directly and indirectly, passing through daylight \((\text{D})\). Additionally, window shading \((\text{W})\) also indirectly influences visual comfort \((\text{V})\) through daylight \((\text{D})\).
Code
dag_coords.ex3 <-
data.frame(name = c('W', 'D', 'V', 'P'),
x = c(1, 3.5, 6, 6),
y = c(3, 2, 1, 3))
DAG.ex3 <-
dagify(D ~ W,
P ~ W + D,
V ~ D,
coords = dag_coords.ex3)
node_labels <- c(
W = 'bold(W)',
D = 'bold(D)',
V = 'bold(V)',
P = 'bold(P)'
)
ggplot(data = DAG.ex3, aes(x = x, y = y, xend = xend, yend = yend)) +
geom_dag_text(aes(label = node_labels[name]),
parse = TRUE,
colour = 'black',
size = 10,
family = 'mono') +
geom_dag_edges(arrow_directed = grid::arrow(length = grid::unit(10, 'pt'), type = 'open'),
edge_colour = 'black',
family = 'mono',
fontface = 'bold') +
annotate('text', x = 1, y = 3.3, label = 'window shading',
size = 4, hjust = 0.5, colour = 'grey50') +
annotate('text', x = 3.5, y = 1.7, label = 'daylight',
size = 4, hjust = 0.6, colour = 'grey50') +
annotate('text', x = 6, y = 3.3, label = 'perceived warmth',
size = 4, hjust = 0.6, colour = 'grey50') +
annotate('text', x = 6, y = 0.7, label = 'visual comfort',
size = 4, hjust = 0.6, colour = 'grey50') +
coord_cartesian(xlim = c(0.5, 6.5), ylim = c(0.8, 3.2)) +
theme_dag()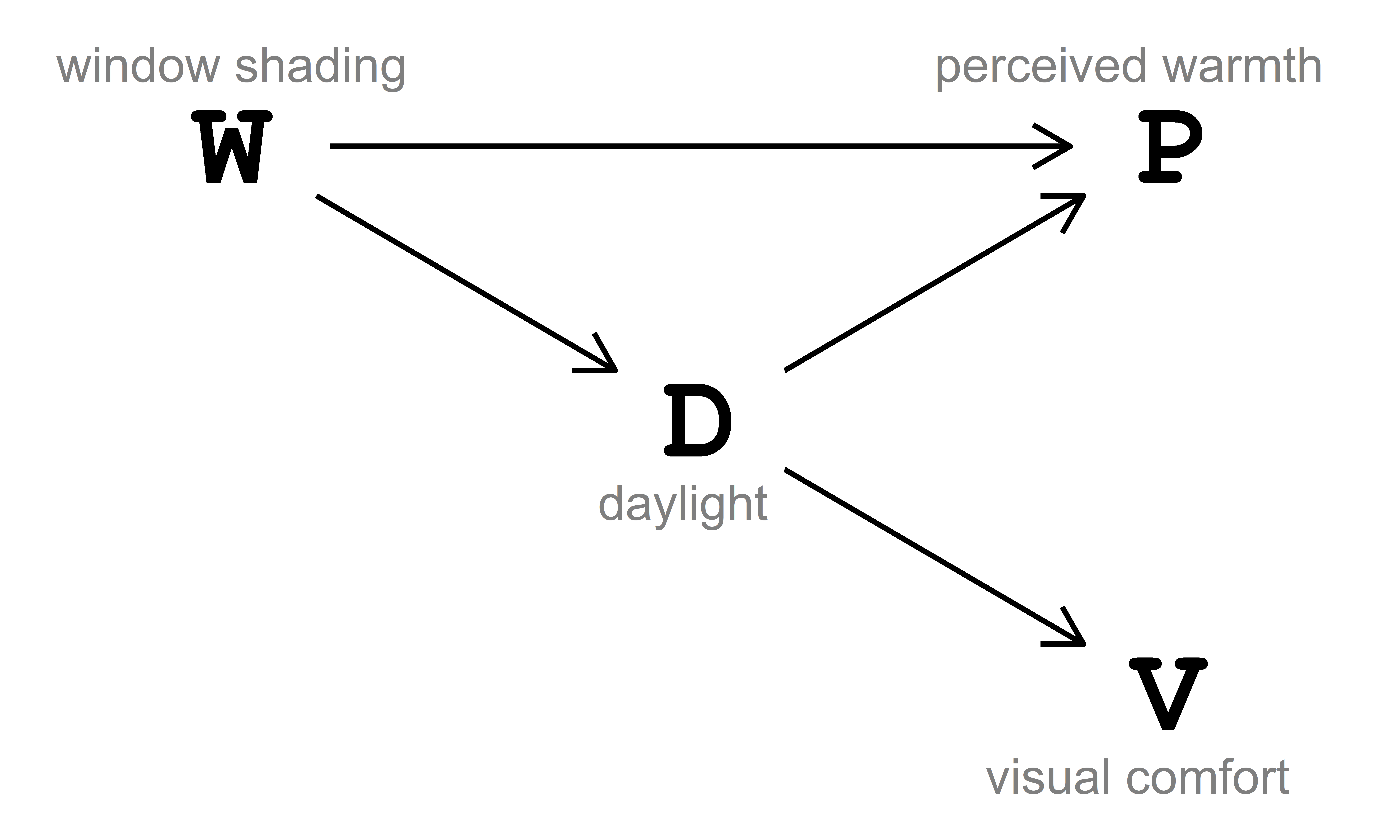
The DAG in Fig. 3.1 can be written as:
- \(D \sim f_{D}(W)\), read as ‘daylight \((\text{D})\) is some function of window shading \((\text{W})\)’.
- \(P \sim f_{P}(W, D)\), read as ‘perceived warmth \((\text{P})\) is some function of window shading \((\text{W})\) and daylight \((\text{D})\)’.
- \(V \sim f_{V}(D)\), read as ‘visual comfort \((\text{V})\) is some function of daylight \((\text{D})\)’.
3.2 Synthetic data set
To generate synthetic data, we defined the custom function data.sim_ex3(). This function takes as inputs the sample size n and generates synthetic data according to the DAG in Fig. 3.1.
data.sim_ex3 <- function(n) {
b_win.day = 1.5 #direct causal effect of W on D
b_win.per = -0.7 #direct causal effect of W on P
b_day.per = 0.8 #direct causal effect of D on P
b_win = b_win.per + (b_win.day * b_day.per) #total causal effect of W on P
#Simulate window
W <- rnorm(n = n, mean = 0, sd = 1)
#Simulate daylight
D <- rnorm(n = n, mean = b_win.day * W, sd = 0.5)
#Simulate visual comfort
V <- rnorm(n = n, mean = 1 * D, sd = 0.2)
#Simulate perceived warmth
P <- rnorm(n = n, mean = b_win.per * W + b_day.per * D, sd = 0.5)
#Return tibble with simulated values
return(tibble(W, D, P, V))
}From this data generation mechanism, we simulated the target population, which consists of one million observations.
set.seed(2025) #set random number for reproducibility
#Simulate the population
ex3_population <- data.sim_ex3(n = 1e6)
#View the data frame
ex3_population# A tibble: 1,000,000 × 4
W D P V
<dbl> <dbl> <dbl> <dbl>
1 0.621 2.01 1.14 1.85
2 0.0356 1.24 1.62 0.875
3 0.773 0.826 -0.716 1.06
4 1.27 0.978 0.241 1.05
5 0.371 0.0871 -0.0980 -0.384
6 -0.163 -0.0222 -0.364 0.387
7 0.397 0.672 0.821 0.867
8 -0.0800 -0.801 -1.27 -0.553
9 -0.345 0.282 1.40 0.280
10 0.702 0.306 -0.193 0.538
# ℹ 999,990 more rowsFrom this population, we obtained one data set of five thousand observations using simple random sampling.
set.seed(ex3_random.seed[1])
#Sample one data set of 5,000 observations
ex3_sample.random <-
ex3_population %>%
slice_sample(n = 5e3) #take a simple random sample of size 5,000
#View the data frame
ex3_sample.random# A tibble: 5,000 × 4
W D P V
<dbl> <dbl> <dbl> <dbl>
1 0.148 0.878 0.767 0.889
2 0.820 1.45 0.348 1.21
3 -1.28 -2.24 -0.939 -2.46
4 -0.179 -0.226 -0.331 -0.352
5 -0.131 -0.320 -0.189 -0.284
6 -0.990 -1.88 -0.432 -1.78
7 -0.522 -1.16 -0.688 -1.53
8 -0.00594 -0.293 0.398 -0.335
9 -0.146 -0.950 0.0766 -0.574
10 -0.991 -2.15 -1.50 -2.12
# ℹ 4,990 more rows3.3 Data analysis
In this example, the target of our analysis is the total average causal effect, ACE (also known as total average treatment effect, ATE) of window shading \((\text{W})\) on perceived warmth \((\text{P})\), which stands for the expected increase of \(\text{P}\) in response to a unit increase in \(\text{W}\) due to an intervention. The causal effect of interest is visualized in Fig. 3.2.
Code
ggplot(data = DAG.ex3, aes(x = x, y = y, xend = xend, yend = yend)) +
#visualize causal effect path
geom_segment(x = 1, xend = 6, y = 3, yend = 3,
linewidth = 14, lineend = 'round', colour = '#009E73', alpha = 0.05) +
geom_segment(x = 1, xend = 3.5, y = 3, yend = 2,
linewidth = 14, lineend = 'round', colour = '#009E73', alpha = 0.05) +
geom_segment(x = 3.5, xend = 6, y = 2, yend = 3,
linewidth = 14, lineend = 'round', colour = '#009E73', alpha = 0.05) +
geom_dag_text(aes(label = node_labels[name]),
parse = TRUE,
colour = 'black',
size = 10,
family = 'mono') +
geom_dag_edges(arrow_directed = grid::arrow(length = grid::unit(10, 'pt'), type = 'open'),
edge_colour = 'black',
family = 'mono',
fontface = 'bold') +
annotate('text', x = 1, y = 3.3, label = 'window shading',
size = 4, hjust = 0.5, colour = 'grey50') +
annotate('text', x = 3.5, y = 1.7, label = 'daylight',
size = 4, hjust = 0.6, colour = 'grey50') +
annotate('text', x = 6, y = 3.3, label = 'perceived warmth',
size = 4, hjust = 0.6, colour = 'grey50') +
annotate('text', x = 6, y = 0.7, label = 'visual comfort',
size = 4, hjust = 0.6, colour = 'grey50') +
#causal effect number
annotate('text', x = 3.5, y = 3.2, label = '(-0.7)',
size = 4.5, hjust = 0.5, colour = 'grey50', parse = TRUE) +
annotate('text', x = 1.8, y = 2.4, label = '(1.5)',
size = 4.5, hjust = 0.5, colour = 'grey50', parse = TRUE) +
annotate('text', x = 5.2, y = 2.4, label = '(0.8)',
size = 4.5, hjust = 0.5, colour = 'grey50', parse = TRUE) +
#causal effect total number b_win = b_win.per + (b_win.day * b_day.per)
annotate('text', x = 3.5, y = 2.8, label = '-0.7 + (1.5 %*% 0.8) == 0.5', # %->%
size = 4.5, hjust = 0.5, colour = 'black', parse = TRUE) +
coord_cartesian(xlim = c(0.5, 6.5), ylim = c(0.8, 3.2)) +
theme_dag()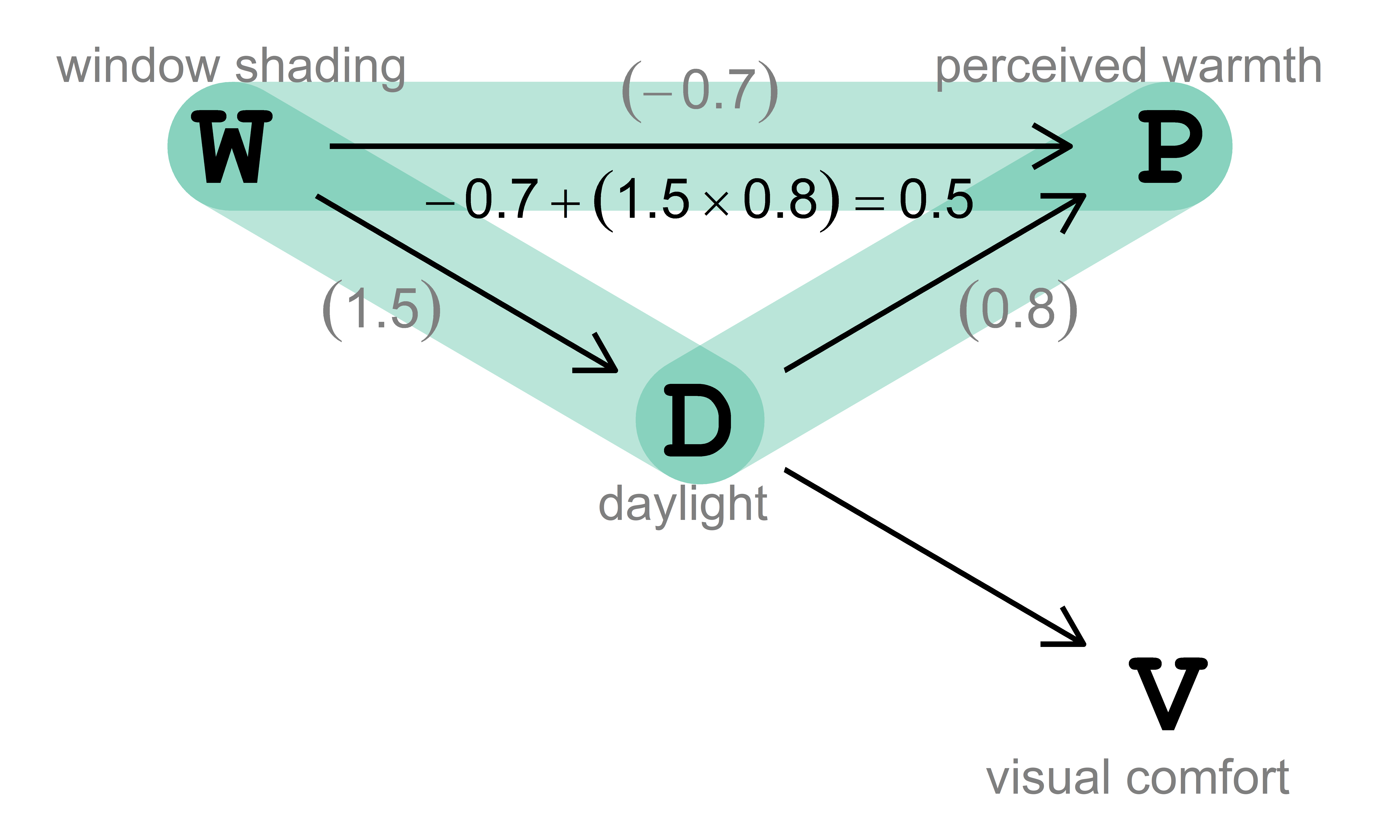
3.3.1 Identification
The first step to answer the causal question of interest is identification. Identification answers a ‘theoretical’ question by determining whether a causal effect can, in principle, be estimated from observed data. The backdoor criterion and its generalization, the adjustment criterion, allow us to understand whether our causal effect of interest can be identified and, if so, which variables we should (or should not) statistically adjust for (i.e., the adjustment set) to estimate the causal effect from the data.
Given its simplicity, we will first apply the backdoor criterion to identify valid adjustment sets to estimate the causal effect of interest. If the backdoor criterion is not applicable, we will apply its generalization, the adjustment criterion.
Backdoor criterion
Applying the backdoor criterion revealed the absence of any backdoor path (i.e., a non-causal path) from window shading \((\text{W})\) to perceived warmth \((\text{P})\). As a result, there is no confounding and no adjustment is required.
Given the DAG in Fig. 3.2, we can use the adjustmentSets() function to identify the adjustment set algorithmically. It is essential to note that this function applies the adjustment criterion and not the backdoor criterion. As such, the adjustmentSets() function can find adjustment sets even when the backdoor criterion is not applicable.
adjustmentSets(DAG.ex3,
exposure = 'W', #window shading
outcome = 'P', #perceived warmth
type = 'all',
effect = 'total',
max.results = Inf)As expected, the resulting adjustment set includes the empty set. Therefore, to get the correct estimate of the total average causal effect, no adjustment is required; failing to do so will lead to bias.
3.3.2 Estimation
Following the identification step is the estimation step. This step addresses a statistical question by determining how the causal effect identified in the previous step can be estimated. To perform this step, we used a parametric (model-based) estimator, specifically, linear regression. This was possible because we designed the illustrative examples to be simple and with a linear relationship between the variables. This way, we limited the complexity of the examples themselves and shifted the focus to the application of the backdoor criterion to define ‘correct’ adjustment sets.
For transparency and understanding, all (implicit) assumptions used for this illustrative example are (explicitly) provided in Table 3.1.
| Research question | Total average causal effect (ACE) of window shading (W) on perceived warmth (P). |
| Assumptions | Random sample (simple random sampling): everyone in the population has an equal chance of being selected into the sample. |
| Limited random variability: large sample size. | |
| Independence of observations: each observation represents independent bits of information. | |
| No confounding: the DAG includes all shared causes among the variables. | |
| No model error: perfect functional form specification. | |
| No measurement error: all variables are measured perfectly. | |
| Variables | window shading (W): continuous variable [unit: -] |
| Daylight (D): continuous variable [unit: lux] | |
| Perceived warmth (P): continuous variable [unit: -] | |
| Visual comfort (V): continuous variable [unit: -] |
To carry out the estimation step, we utilized linear regression within both the frequentist and Bayesian frameworks. Specifically, we will run three regression models:
-
Model.1will include only window shading \((\text{W})\) as predictor; -
Model.2will include window shading \((\text{W})\) and daylight \((\text{D})\) as predictors. -
Model.3will include window shading \((\text{W})\) and visual comfort \((\text{V})\) as predictors.
The results of the fitted statistical models (i.e., Model.1, Model.2 and Model.3) are presented here.
Frequentist framework
#Fit the linear regression model with W and P (Model 1)
ex3_Model.1 <-
lm(formula = P ~ W,
data = ex3_sample.random)
#View of the model summary
summary(ex3_Model.1)
Call:
lm(formula = P ~ W, data = ex3_sample.random)
Residuals:
Min 1Q Median 3Q Max
-2.73804 -0.42710 0.00262 0.42714 2.35619
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.001521 0.009025 -0.169 0.866
W 0.510180 0.008920 57.192 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.6382 on 4998 degrees of freedom
Multiple R-squared: 0.3956, Adjusted R-squared: 0.3954
F-statistic: 3271 on 1 and 4998 DF, p-value: < 2.2e-16#Fit the linear regression model with W, D and P (Model 2)
ex3_Model.2 <-
lm(formula = P ~ W + D,
data = ex3_sample.random)
#View of the model summary
summary(ex3_Model.2)
Call:
lm(formula = P ~ W + D, data = ex3_sample.random)
Residuals:
Min 1Q Median 3Q Max
-1.92355 -0.33391 -0.00018 0.33940 1.68471
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.003004 0.007111 0.422 0.673
W -0.698074 0.022967 -30.394 <2e-16 ***
D 0.800956 0.014495 55.259 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.5028 on 4997 degrees of freedom
Multiple R-squared: 0.6248, Adjusted R-squared: 0.6247
F-statistic: 4161 on 2 and 4997 DF, p-value: < 2.2e-16#Fit the linear regression model with W, V and P (Model 3)
ex3_Model.3 <-
lm(formula = P ~ W + V,
data = ex3_sample.random)
#View of the model summary
summary(ex3_Model.3)
Call:
lm(formula = P ~ W + V, data = ex3_sample.random)
Residuals:
Min 1Q Median 3Q Max
-1.9264 -0.3466 -0.0032 0.3518 1.8682
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.005649 0.007426 0.761 0.447
W -0.522315 0.022364 -23.355 <2e-16 ***
V 0.683934 0.013994 48.873 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.525 on 4997 degrees of freedom
Multiple R-squared: 0.591, Adjusted R-squared: 0.5909
F-statistic: 3611 on 2 and 4997 DF, p-value: < 2.2e-16The estimated coefficients for the three models are then plotted in Fig. 3.3.
Code
b_win.day = 1.5 #direct causal effect of W on D
b_win.per = -0.7 #direct causal effect of W on P
b_day.per = 0.8 #direct causal effect of D on P
b_win = b_win.per + (b_win.day * b_day.per) #total causal effect of W on P
data.frame(model = c('Model.1', 'Model.2', 'Model.3'),
estimate = c(coef(ex3_Model.1)['W'],
coef(ex3_Model.2)['W'],
coef(ex3_Model.3)['W']),
lower.95.CI = c(confint(ex3_Model.1, level = 0.95, type = 'Wald')['W', 1],
confint(ex3_Model.2, level = 0.95, type = 'Wald')['W', 1],
confint(ex3_Model.3, level = 0.95, type = 'Wald')['W', 1]),
upper.95.CI = c(confint(ex3_Model.1, level = 0.95, type = 'Wald')['W', 2],
confint(ex3_Model.2, level = 0.95, type = 'Wald')['W', 2],
confint(ex3_Model.3, level = 0.95, type = 'Wald')['W', 2])) %>%
ggplot(aes(x = estimate, y = model, xmin = lower.95.CI, xmax = upper.95.CI)) +
geom_vline(xintercept = b_win, alpha = 0.8, linetype = 'dashed', colour = 'blue') +
geom_linerange() +
geom_point(shape = 21, size = 2, fill = 'white', stroke = 1) +
scale_x_continuous('Estimate',
breaks = seq(from = -5, to = 5, by = 0.25),
limits = c(-0.85, 0.55)) +
theme(panel.grid = element_blank(),
panel.background = element_blank(),
panel.border = element_rect(colour = 'black', fill = NA),
axis.title.y = element_blank())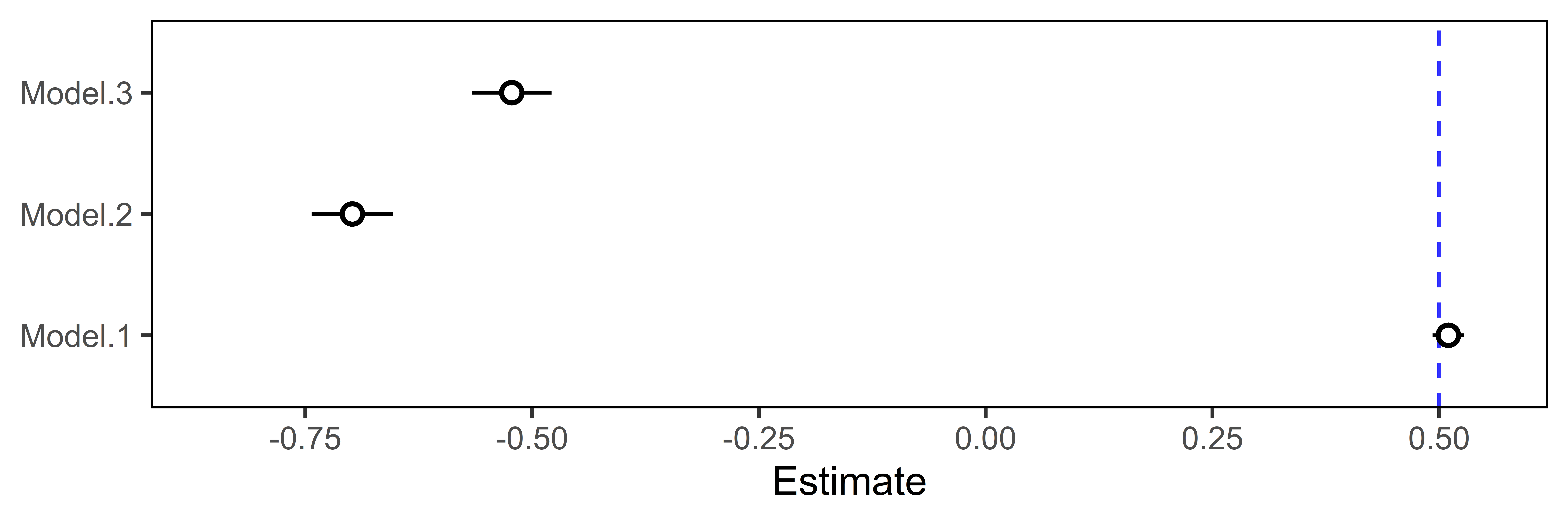
Fig. 3.3 shows the estimates (point estimate and 95% confidence interval) of the coefficient for window shading for Model.1, Model.2 and Model.3.
For Model.1 we found a positive coefficient between window shading \((\text{W})\) and perceived warmth \((\text{P})\) equal to 0.510 with 95% confidence interval (CI) [0.493, 0.528]. Since the 95% CI excludes zero, the regression coefficient is statistically significantly different from zero at the 0.05 level (p-value = 0). Additionally, since the 95% CI includes the data-generating parameter for window shading (i.e., b_win = 0.5), we can deduce that the estimated coefficient for window shading is not statistically significantly different from 0.5 at the 0.05 level (although this will be the case for all numbers within the 95% confidence interval). We can test this more formally by using the linearHypothesis() function. Specifically,
#Test for W = 0.5
car::linearHypothesis(ex3_Model.1, 'W = 0.5') # car:: access the function from the car package without loading it in the environment
Linear hypothesis test:
W = 0.5
Model 1: restricted model
Model 2: P ~ W
Res.Df RSS Df Sum of Sq F Pr(>F)
1 4999 2035.9
2 4998 2035.4 1 0.53037 1.3024 0.2538The resulting p-value is 0.254, which indicates that we fail to reject the null hypothesis (i.e., W = 0.5) at the 0.05 level. This result suggests that the regression coefficient for window shading (i.e., 0.510) is not statistically significantly different from 0.5. Since the estimated causal effect from Model.1 is unbiased, we would correctly conclude that an increase of unit in the opening of window shading causes an increase in perceived warmth by 0.510 units (95% CI [0.493, 0.528]).
For Model.2 we found a negative coefficient between window shading \((\text{W})\) and perceived warmth \((\text{P})\) equal to -0.698 with 95% CI [-0.743, -0.653]. Since the 95% CI excludes zero, the regression coefficient is statistically significantly different from zero at the 0.05 level (p-value = 2.43e-186). Additionally, since the 95% CI does not include the data-generating parameter for window shading (i.e., b_win = 0.5), we can deduce that the estimated coefficient for window shading is statistically significantly different from 0.5 at the 0.05 level. We can test this more formally by using the linearHypothesis() function. Specifically,
#Test for W = 0.5
car::linearHypothesis(ex3_Model.2, 'W = 0.5') # car:: access the function from the car package without loading it in the environment
Linear hypothesis test:
W = 0.5
Model 1: restricted model
Model 2: P ~ W + D
Res.Df RSS Df Sum of Sq F Pr(>F)
1 4998 1951.3
2 4997 1263.4 1 687.97 2721.1 < 2.2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1The resulting p-value is 0, which indicates that we can reject the null hypothesis (i.e., W = 0.5) at the 0.05 level. This result suggests that the regression coefficient for window shading (i.e., -0.698) is statistically significantly different from 0.5. Since the estimated causal effect from Model.2 is biased, we would erroneously conclude that an increase of unit in the opening of window shading causes a decrease in perceived warmth by -0.698 units (95% CI [-0.743, -0.653]).
For Model.3 we found a negative coefficient between window shading \((\text{W})\) and perceived warmth \((\text{P})\) equal to -0.522 with 95% CI [-0.566, -0.478]. Since the 95% CI excludes zero, the regression coefficient is statistically significantly different from zero at the 0.05 level (p-value = 1.39e-114). Additionally, since the 95% CI does not include the data-generating parameter for window shading (i.e., b_win = 0.5), we can deduce that the estimated coefficient for window shading is statistically significantly different from 0.5 at the 0.05 level. We can test this more formally by using the linearHypothesis() function. Specifically,
#Test for W = 0.5
car::linearHypothesis(ex3_Model.3, 'W = 0.5') # car:: access the function from the car package without loading it in the environment
Linear hypothesis test:
W = 0.5
Model 1: restricted model
Model 2: P ~ W + V
Res.Df RSS Df Sum of Sq F Pr(>F)
1 4998 1953.0
2 4997 1377.1 1 575.86 2089.6 < 2.2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1The resulting p-value is 0, which indicates that we can reject the null hypothesis (i.e., W = 0.5) at the 0.05 level. This result suggests that the regression coefficient for window shading (i.e., -0.522) is statistically significantly different from 0.5. Since the estimated causal effect from Model.3 is biased, we would erroneously conclude that an increase of unit in the opening of window shading causes a decrease in perceived warmth by -0.522 units (95% CI [-0.566, -0.478]).
Importantly, for Model.1, Model.2, and Model.3, the 95% confidence interval means that if we were to repeat the sampling process and calculate the interval many times, 95% of those calculated intervals would contain the true population parameter. To highlight this, we can repeat the analysis by fitting the three models to one thousand data sets randomly selected from our population.
The for-loop shown in the code below performs the following operations. First, sample (using simple random sampling) a data set of 5,000 observations from the target population. Subsequently, perform linear regression using Model.1, Model.2 and Model.3 and store the estimated coefficients for window shading, its standard error and 95% confidence interval in the data frame coefs_ex3. This operation is repeated a thousand times, resulting in the data frame coefs_ex3 containing the estimates (point estimate, standard error and confidence interval) of a thousand random samples of size 5,000 using Model.1, Model.2 and Model.3.
n_model <- c('mod.1', 'mod.2', 'mod.3')
n_row <- n_sims*length(n_model)
#Create an empty data frame
empty.df <- data.frame(matrix(NA, nrow = n_row, ncol = 7))
#Rename the data frame columns
colnames(empty.df) <- c('sim.id', 'estimate', 'se', 'CI_2.5', 'CI_97.5',
'model', 'coverage')
#Sample a thousand data sets of 5,000 observations and perform linear regression
coefs_ex3 <- empty.df #assign the empty data frame
k = 1
for (i in 1:n_sims){
set.seed(ex3_random.seed[i]) #set unique seed for each simulation
#Sample data set from population
sample.random <-
ex3_population %>%
slice_sample(n = 5e3) #take a simple random sample of size 5,000
#Fit models
for (j in 1:length(n_model)){
if (n_model[j] == 'mod.1'){
fit <- lm(formula = P ~ W,
data = sample.random)
} else if (n_model[j] == 'mod.2'){
fit <- lm(formula = P ~ W + D,
data = sample.random)
} else {
fit <- lm(formula = P ~ W + V,
data = sample.random)
}
#Compile matrix
coefs_ex3[k, 1] <- i #simulation ID
coefs_ex3[k, 2] <- coef(fit)['W'] #point estimate
coefs_ex3[k, 3] <- summary(fit)$coef['W','Std. Error'] #standard error
coefs_ex3[k, 4:5] <- confint(fit, level = 0.95, type = 'Wald')['W', ] #confidence interval (Wald)
coefs_ex3[k, 6] <- n_model[j] #sample size
k = k + 1
}
}
coefs_ex3 <- as_tibble(coefs_ex3)
#View the data frame
coefs_ex3# A tibble: 3,000 × 7
sim.id estimate se CI_2.5 CI_97.5 model coverage
<int> <dbl> <dbl> <dbl> <dbl> <chr> <lgl>
1 1 0.510 0.00892 0.493 0.528 mod.1 NA
2 1 -0.698 0.0230 -0.743 -0.653 mod.2 NA
3 1 -0.522 0.0224 -0.566 -0.478 mod.3 NA
4 2 0.488 0.00913 0.470 0.506 mod.1 NA
5 2 -0.710 0.0221 -0.753 -0.666 mod.2 NA
6 2 -0.555 0.0216 -0.597 -0.512 mod.3 NA
7 3 0.495 0.00903 0.478 0.513 mod.1 NA
8 3 -0.707 0.0220 -0.750 -0.664 mod.2 NA
9 3 -0.544 0.0216 -0.586 -0.502 mod.3 NA
10 4 0.511 0.00910 0.494 0.529 mod.1 NA
# ℹ 2,990 more rowsThe coverage is defined by setting its value to 1 if the confidence interval overlaps the data-generating parameter for window shading (i.e., b_win = 0.5) and 0 otherwise.
#Calculate coverage
coefs_ex3 <-
coefs_ex3 %>%
mutate(coverage = case_when(CI_2.5 > b_win | CI_97.5 < b_win ~ 0,
CI_2.5 <= b_win & CI_97.5 >= b_win ~ 1,
.default = NA))
#View the data frame
coefs_ex3# A tibble: 3,000 × 7
sim.id estimate se CI_2.5 CI_97.5 model coverage
<int> <dbl> <dbl> <dbl> <dbl> <chr> <dbl>
1 1 0.510 0.00892 0.493 0.528 mod.1 1
2 1 -0.698 0.0230 -0.743 -0.653 mod.2 0
3 1 -0.522 0.0224 -0.566 -0.478 mod.3 0
4 2 0.488 0.00913 0.470 0.506 mod.1 1
5 2 -0.710 0.0221 -0.753 -0.666 mod.2 0
6 2 -0.555 0.0216 -0.597 -0.512 mod.3 0
7 3 0.495 0.00903 0.478 0.513 mod.1 1
8 3 -0.707 0.0220 -0.750 -0.664 mod.2 0
9 3 -0.544 0.0216 -0.586 -0.502 mod.3 0
10 4 0.511 0.00910 0.494 0.529 mod.1 1
# ℹ 2,990 more rowsThe results are then plotted in Fig. 3.4.
Code
model_names <- c('mod.1' = 'Model.1',
'mod.2' = 'Model.2',
'mod.3' = 'Model.3')
ggplot(data = subset(coefs_ex3, sim.id <= 2e2), aes(x = sim.id, y = estimate, ymin = CI_2.5, ymax = CI_97.5, colour = as.factor(coverage))) +
geom_hline(yintercept = b_win, alpha = 0.8, linetype = 'dashed', colour = 'blue') +
geom_linerange() +
geom_point(shape = 21, size = 1, fill = 'white', stroke = 0.5) +
scale_colour_manual('', values = c('black', '#FF4040'),
breaks = c('1','0')) +
scale_y_continuous('Estimate') +
scale_x_continuous('Simulation ID') +
facet_wrap(model~.,
labeller = labeller(model = model_names),
nrow = 3,
scales = 'fixed') +
theme(legend.position = 'none',
panel.grid = element_blank(),
panel.background = element_blank(),
panel.border = element_rect(colour = 'black', fill = NA),
axis.title.y = element_blank())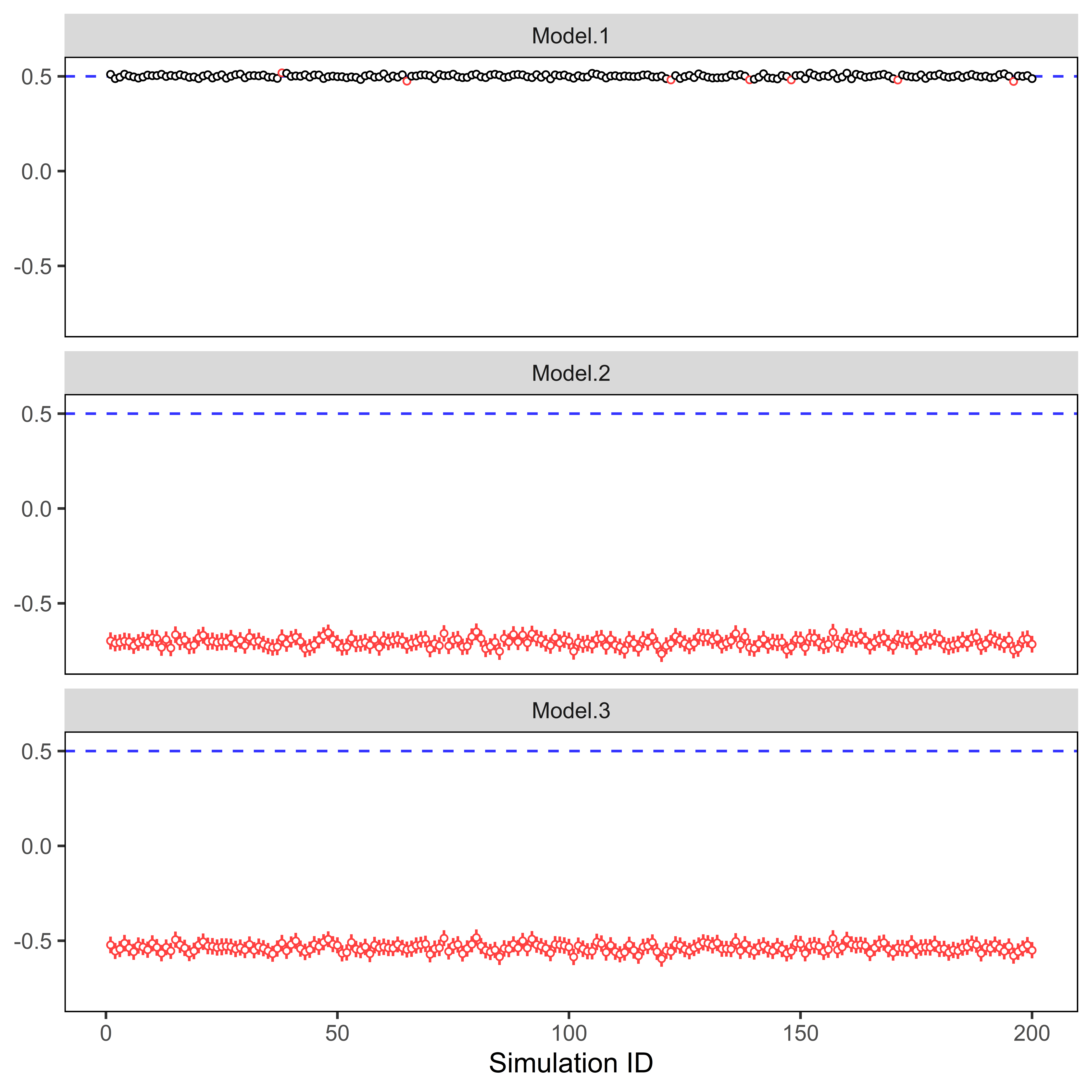
Fig. 3.4 shows the first 200 estimates (point estimate and confidence interval) of the causal effect of window shading for Model.1, Model.2 and Model.3. If all the thousand simulations are considered, the frequency of the coverage of the calculated confidence intervals (i.e., how many times the confidence intervals overlap the data-generating parameter) is 94.8%, 0.0% and 0.0% for Model.1, Model.2 and Model.3, respectively. Since the estimate of the causal effect for Model.2 and Model.3 are biased, the calculated confidence intervals for these models do not have the expected coverage. In fact, confidence intervals only quantify the uncertainty due to random error (i.e., sample variability), not systematic error (i.e., bias). Instead, the estimates from Model.1 are unbiased and the calculated 95% confidence intervals have the expected coverage (i.e., overlap the data-generating parameter 95% of the times).
We can also visualize all the 1,000 estimates of the coefficient for window shading for Model.1, Model.2 and Model.3 (the estimates were shown as white dots in Fig. 3.4).
Code
ggplot(data = coefs_ex3, aes(x = estimate, y = ifelse(after_stat(count) > 0, after_stat(count), NA))) +
geom_histogram(binwidth = 0.015, fill = 'white', colour = 'grey50') +
geom_vline(xintercept = b_win, colour = 'blue', linetype = 'dashed') +
scale_x_continuous('Estimate',
breaks = seq(from = -5, to = 5, by = 0.25),
limits = c(-0.85, 0.55)) +
facet_grid(model~.,
labeller = labeller(model = model_names),
scales = 'fixed') +
theme(axis.title.y = element_blank(),
axis.text.y = element_blank(),
axis.ticks.y = element_blank(),
panel.grid = element_blank(),
panel.background = element_blank(),
panel.border = element_rect(colour = 'black', fill = NA))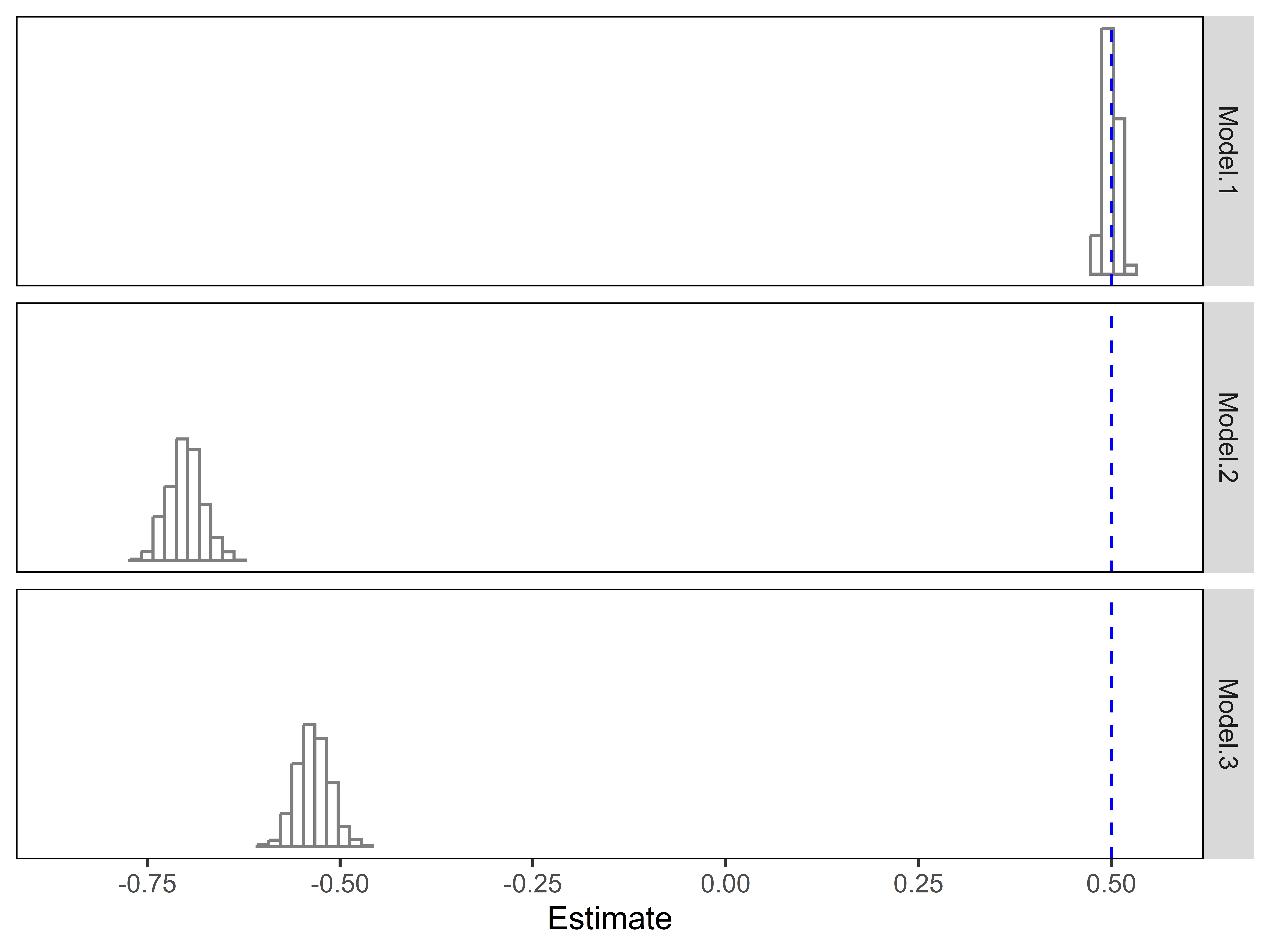
In Fig. 3.5, the estimated coefficients are visualized through a histogram. Here, we can see that the estimates for Model.2 and Model.3 are not clustered around the data-generating parameter (blue dashed line) having a mean estimate of -0.701 (with 0.022 standard deviation) and -0.535 (with 0.021 standard deviation), respectively. In contrast, the estimates for Model.1 having a mean estimate of 0.499 (with 0.009 standard deviation) are centered around the data-generating parameter.
As expected, since there is no backdoor path open, regressing perceived warmth \((\text{P})\) on window shading \((\text{W})\) (i.e., using Model.1) leads to the correct estimate of the total average causal effect. However, when daylight is included as a predictor (i.e., using Model.2), the estimates are biased. Adjusting for daylight blocks a part of the causal effect. This happens because daylight is a mediator on the path from window shading \((\text{W})\) to perceived warmth \((\text{P})\). This bias is known as post-treatment bias. Additionally, we have a biased estimate of the total average causal effect of window shading \((\text{W})\) on perceived warmth \((\text{P})\) when we include visual comfort \((\text{V})\) as a predictor (i.e., using Model.3). This happens because visual comfort \((\text{V})\) is a descendant of daylight \((\text{D})\); consequently, adjusting for visual comfort \((\text{V})\) is akin to adjusting for daylight \((\text{D})\) since the two variables share information (daylight is the cause of visual comfort).
Bayesian framework
Click to expand
#Fit the linear regression model with W and P (Model 1)
ex3_Model.1 <- stan_glm(formula = P ~ W,
family = gaussian(),
data = ex3_sample.random,
#Prior coefficients
prior = normal(location = 0, scale = 1),
#Prior intercept
prior_intercept = normal(location = 0, scale = 1),
#Prior sigma
prior_aux = exponential(rate = 1),
iter = 4000, warmup = 1000,
save_warmup = TRUE,
chains = 4, cores = 4,
seed = 2025) #for reproducibility
#View of the model
ex3_Model.1stan_glm
family: gaussian [identity]
formula: P ~ W
observations: 5000
predictors: 2
------
Median MAD_SD
(Intercept) 0.0 0.0
W 0.5 0.0
Auxiliary parameter(s):
Median MAD_SD
sigma 0.6 0.0
------
* For help interpreting the printed output see ?print.stanreg
* For info on the priors used see ?prior_summary.stanreg#Fit the linear regression model with W, D and P (Model 2)
ex3_Model.2 <- stan_glm(formula = P ~ W + D,
family = gaussian(),
data = ex3_sample.random,
#Prior coefficients
prior = normal(location = c(0, 0), scale = c(1, 1)),
#Prior intercept
prior_intercept = normal(location = 0, scale = 1),
#Prior sigma
prior_aux = exponential(rate = 1),
iter = 4000, warmup = 1000,
save_warmup = TRUE,
chains = 4, cores = 4,
seed = 2025) #for reproducibility
#View of the model
ex3_Model.2stan_glm
family: gaussian [identity]
formula: P ~ W + D
observations: 5000
predictors: 3
------
Median MAD_SD
(Intercept) 0.0 0.0
W -0.7 0.0
D 0.8 0.0
Auxiliary parameter(s):
Median MAD_SD
sigma 0.5 0.0
------
* For help interpreting the printed output see ?print.stanreg
* For info on the priors used see ?prior_summary.stanreg#Fit the linear regression model with W, V and P (Model 3)
ex3_Model.3 <- stan_glm(formula = P ~ W + V,
family = gaussian(),
data = ex3_sample.random,
#Prior coefficients
prior = normal(location = c(0, 0), scale = c(1, 1)),
#Prior intercept
prior_intercept = normal(location = 0, scale = 1),
#Prior sigma
prior_aux = exponential(rate = 1),
iter = 4000, warmup = 1000,
save_warmup = TRUE,
chains = 4, cores = 4,
seed = 2025) #for reproducibility
#View of the model
ex3_Model.3stan_glm
family: gaussian [identity]
formula: P ~ W + V
observations: 5000
predictors: 3
------
Median MAD_SD
(Intercept) 0.0 0.0
W -0.5 0.0
V 0.7 0.0
Auxiliary parameter(s):
Median MAD_SD
sigma 0.5 0.0
------
* For help interpreting the printed output see ?print.stanreg
* For info on the priors used see ?prior_summary.stanregIn Bayesian analysis, there are important diagnostics that have to be carried out in order to assess the convergence and efficiency of the Markov Chains. This is done by using the monitor() function which computes summaries of MCMC (Markov Chain Monte Carlo) draws and monitor convergence. Specifically, we will look at Rhat, Bulk_ESS and Tail_ESS metrics.
#Diagnostics for model 1
monitor(ex3_Model.1$stanfit) Inference for the input samples (4 chains: each with iter = 4000; warmup = 0):
Q5 Q50 Q95 Mean SD Rhat Bulk_ESS Tail_ESS
(Intercept) 0.0 0.0 0.0 0.0 0.0 1 12088 8622
W 0.5 0.5 0.5 0.5 0.0 1 11533 8670
sigma 0.6 0.6 0.6 0.6 0.0 1 11793 8110
mean_PPD 0.0 0.0 0.0 0.0 0.0 1 12191 10616
log-posterior -4854.8 -4852.0 -4851.0 -4852.4 1.2 1 6031 7127
For each parameter, Bulk_ESS and Tail_ESS are crude measures of
effective sample size for bulk and tail quantities respectively (an ESS > 100
per chain is considered good), and Rhat is the potential scale reduction
factor on rank normalized split chains (at convergence, Rhat <= 1.05).#Diagnostics for model 2
monitor(ex3_Model.2$stanfit) Inference for the input samples (4 chains: each with iter = 4000; warmup = 0):
Q5 Q50 Q95 Mean SD Rhat Bulk_ESS Tail_ESS
(Intercept) 0.0 0.0 0.0 0.0 0.0 1 7636 6818
W -0.7 -0.7 -0.7 -0.7 0.0 1 4770 5567
D 0.8 0.8 0.8 0.8 0.0 1 4796 5487
sigma 0.5 0.5 0.5 0.5 0.0 1 8255 6236
mean_PPD 0.0 0.0 0.0 0.0 0.0 1 9358 10287
log-posterior -3664.7 -3661.7 -3660.4 -3662.0 1.4 1 4692 6853
For each parameter, Bulk_ESS and Tail_ESS are crude measures of
effective sample size for bulk and tail quantities respectively (an ESS > 100
per chain is considered good), and Rhat is the potential scale reduction
factor on rank normalized split chains (at convergence, Rhat <= 1.05).#Diagnostics for model 3
monitor(ex3_Model.3$stanfit) Inference for the input samples (4 chains: each with iter = 4000; warmup = 0):
Q5 Q50 Q95 Mean SD Rhat Bulk_ESS Tail_ESS
(Intercept) 0.0 0.0 0.0 0.0 0.0 1 8298 7180
W -0.6 -0.5 -0.5 -0.5 0.0 1 4298 5085
V 0.7 0.7 0.7 0.7 0.0 1 4334 5399
sigma 0.5 0.5 0.5 0.5 0.0 1 9060 7011
mean_PPD 0.0 0.0 0.0 0.0 0.0 1 9705 10021
log-posterior -3880.2 -3877.1 -3875.7 -3877.4 1.4 1 4575 6066
For each parameter, Bulk_ESS and Tail_ESS are crude measures of
effective sample size for bulk and tail quantities respectively (an ESS > 100
per chain is considered good), and Rhat is the potential scale reduction
factor on rank normalized split chains (at convergence, Rhat <= 1.05).Rhat is a metric used to assess the convergence of Markov Chain Monte Carlo (MCMC) simulations. It helps determine if the MCMC chains have adequately explored the target posterior distribution. Specifically, it compares the between- and within-chain estimates for model parameters: If chains have not mixed well (i.e., the between- and within-chain estimates do not agree), R-hat is larger than 1. A general rule of thumb is to use the sample only if R-hat is less than 1.05; a larger value suggests that the chains have not mixed well, and the results might not be reliable. In our three models, all Rhat are equal to 1 indicating that the chains have mixed well and have adequately explored the target posterior distribution.
Bulk_ESS and Tail_ESS stand for ‘Bulk Effective Sample Size’ and ‘Tail Effective Sample Size,’ respectively. Since MCMC samples are not truly independent (they are correlated), these metrics assess the sampling efficiencies, that is, they help evaluate how efficiently the MCMC sampler is exploring the parameter space.
-
Bulk_ESSis a useful measure for sampling efficiency in the bulk (center) of the distribution (e.g., efficiency of mean and median estimates); -
Tail_ESSis a useful measure for sampling efficiency in the tails of the distribution (e.g., efficiency of variance and tail quantile estimates). A general rule of thumb is that bothBulk-ESSandTail-ESSshould be at least 100 (approximately) per Markov Chain in order to be reliable and indicate that estimates of respective posterior quantiles are reliable. In our three models, allBulk-ESSandTail-ESSare well above 400 (i.e., 100 multiplied by 4, the number of chains we used) indicating that estimates of posterior quantiles are reliable.
Since we have established that the posteriors are reliable, we can now explore the model estimates.
The estimated coefficients for the three models are then plotted in Fig. 3.6.
Code
#Extract draws from model 1
post_ex3_Model.1 <-
ex3_Model.1 %>%
spread_draws(W) %>% #extract draws from the fitted model
mutate(model = 'Model.1') #add a new column to specify that the model
#Extract draws from model 2
post_ex3_Model.2 <-
ex3_Model.2 %>%
spread_draws(W) %>% #extract draws from the fitted model
mutate(model = 'Model.2') #add a new column to specify that the model
#Extract draws from model 3
post_ex3_Model.3 <-
ex3_Model.3 %>%
spread_draws(W) %>% #extract draws from the fitted model
mutate(model = 'Model.3') #add a new column to specify that the model
#Combine draws
plot.post <- rbind(post_ex3_Model.1, post_ex3_Model.2, post_ex3_Model.3)
# Plot
plot.post %>%
ggplot(aes(y = model, x = W)) +
stat_slabinterval(point_interval = 'mean_hdi',
.width = c(.95)) +
geom_vline(xintercept = b_win, alpha = 0.8, linetype = 'dashed', colour = 'blue') +
scale_x_continuous('Estimate',
breaks = seq(from = -5, to = 5, by = 0.25),
limits = c(-0.85, 0.55)) +
theme(panel.grid = element_blank(),
panel.background = element_blank(),
panel.border = element_rect(colour = 'black', fill = NA),
axis.title.y = element_blank())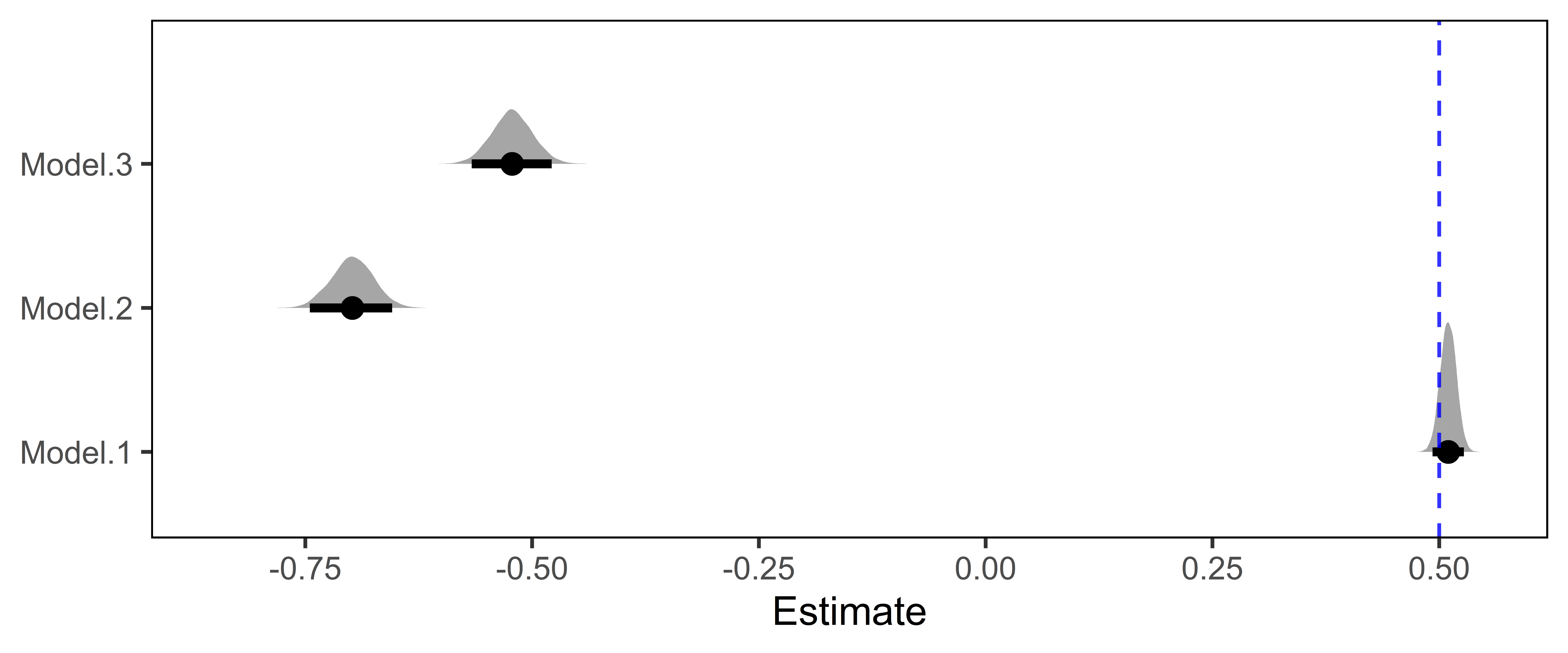
Fig. 3.6 shows the estimates (mean and 95% HDI) of the coefficient for window shading for Model.1, Model.2 and Model.3.
For Model.1 we found a positive coefficient between window shading \((\text{W})\) and perceived warmth \((\text{P})\) (mean = 0.510, 95% HDI [0.493, 0.527]). The estimated causal effect is unbiased, leading to the correct conclusion that an increase of unit in the opening of window shading causes an increase in perceived warmth by 0.510 units, on average.
For Model.2 we found a negative coefficient between window shading \((\text{W})\) and perceived warmth \((\text{P})\) (mean = -0.698, 95% HDI [-0.745, -0.654]). The estimated causal effect is therefore biased, leading to the wrong conclusion that an increase of unit in the opening of window shading causes a decrease in perceived warmth by -0.698 units, on average.
For Model.3 we found a negative coefficient between window shading \((\text{W})\) and perceived warmth \((\text{P})\) (mean = -0.522, 95% HDI [-0.567, -0.478]). The estimated causal effect is therefore biased, leading to the wrong conclusion that an increase of unit in the opening of window shading causes a decrease in perceived warmth by -0.522 units, on average.
Importantly, for both Model.1, Model.2 and Model.3, the 95% HDI is the range of parameter values within which the most credible 95% of the posterior distribution falls. Unlike a frequentist confidence interval, the Bayesian 95% HDI has a direct probabilistic meaning: every point inside the HDI has a higher probability density than any point outside the interval. Therefore, given the model, the prior and the data, we can say that there is a 95% probability that the data-generating parameter (i.e., b_win = 0.5) lies within the HDI. However, since Model.2 and Model.3 lead to biased estimates, we will reach the wrong conclusion by stating that there is a 95% probability that the data-generating parameter lies within the [-0.745, -0.654] interval and [-0.567, -0.478], respectively. This probability is 0%.
As expected, since there is no backdoor path open, regressing perceived warmth \((\text{P})\) on window shading \((\text{W})\) (i.e., using Model.1) leads to the correct estimate of the total average causal effect. However, when daylight is included as a predictor (i.e., using Model.2) the estimates are biased. Adjusting for daylight blocks a part of the causal effect. This happens because daylight is a mediator on the path from window shading \((\text{W})\) to perceived warmth \((\text{P})\). This bias is known as post-treatment bias. Additionally, we have a biased estimate of the total average causal effect of window shading \((\text{W})\) on perceived warmth \((\text{P})\) when we include visual comfort \((\text{V})\) as a predictor (i.e., using Model.3). This happens because visual comfort \((\text{V})\) is a descendant of daylight \((\text{D})\); consequently, adjusting for visual comfort \((\text{V})\) is akin to adjusting for daylight \((\text{D})\) since the two variables share information (daylight is the cause of visual comfort).
Session info
Version information about R, the OS and attached or loaded packages.
R version 4.2.3 (2023-03-15 ucrt)
Platform: x86_64-w64-mingw32/x64 (64-bit)
Running under: Windows 10 x64 (build 26200)
Matrix products: default
locale:
[1] LC_COLLATE=English_United Kingdom.utf8
[2] LC_CTYPE=English_United Kingdom.utf8
[3] LC_MONETARY=English_United Kingdom.utf8
[4] LC_NUMERIC=C
[5] LC_TIME=English_United Kingdom.utf8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] rstan_2.32.7 StanHeaders_2.32.10 gt_1.1.0
[4] lubridate_1.9.4 forcats_1.0.1 stringr_1.6.0
[7] dplyr_1.1.4 purrr_1.2.0 readr_2.1.6
[10] tidyr_1.3.1 tibble_3.3.0 ggplot2_4.0.1
[13] tidyverse_2.0.0 ggdist_3.3.3 tidybayes_3.0.7
[16] rstanarm_2.32.2 Rcpp_1.1.0 dagitty_0.3-4
[19] ggdag_0.2.13
loaded via a namespace (and not attached):
[1] backports_1.5.0 plyr_1.8.9 igraph_2.2.1
[4] splines_4.2.3 svUnit_1.0.8 crosstalk_1.2.2
[7] rstantools_2.5.0 inline_0.3.21 digest_0.6.38
[10] htmltools_0.5.8.1 viridis_0.6.5 magrittr_2.0.4
[13] checkmate_2.3.3 memoise_2.0.1 tzdb_0.5.0
[16] graphlayouts_1.2.2 RcppParallel_5.1.11-1 matrixStats_1.5.0
[19] xts_0.14.1 timechange_0.3.0 colorspace_2.1-2
[22] ggrepel_0.9.6 rbibutils_2.4 xfun_0.54
[25] jsonlite_2.0.0 lme4_1.1-37 survival_3.8-3
[28] zoo_1.8-14 glue_1.8.0 reformulas_0.4.2
[31] polyclip_1.10-7 gtable_0.3.6 V8_8.0.1
[34] distributional_0.5.0 car_3.1-3 pkgbuild_1.4.8
[37] abind_1.4-8 scales_1.4.0 miniUI_0.1.2
[40] viridisLite_0.4.2 xtable_1.8-4 Formula_1.2-5
[43] stats4_4.2.3 DT_0.34.0 htmlwidgets_1.6.4
[46] threejs_0.3.4 arrayhelpers_1.1-0 RColorBrewer_1.1-3
[49] posterior_1.6.1 pkgconfig_2.0.3 loo_2.8.0
[52] farver_2.1.2 sass_0.4.10 utf8_1.2.6
[55] tidyselect_1.2.1 labeling_0.4.3 rlang_1.1.6
[58] reshape2_1.4.5 later_1.4.4 tools_4.2.3
[61] cachem_1.1.0 cli_3.6.5 generics_0.1.4
[64] evaluate_1.0.5 fastmap_1.2.0 yaml_2.3.10
[67] knitr_1.50 fs_1.6.6 tidygraph_1.3.1
[70] ggraph_2.2.2 nlme_3.1-168 mime_0.13
[73] xml2_1.5.0 compiler_4.2.3 bayesplot_1.14.0
[76] shinythemes_1.2.0 rstudioapi_0.17.1 curl_7.0.0
[79] tweenr_2.0.3 stringi_1.8.7 lattice_0.22-7
[82] Matrix_1.6-5 nloptr_2.2.1 markdown_2.0
[85] shinyjs_2.1.0 tensorA_0.36.2.1 vctrs_0.6.5
[88] pillar_1.11.1 lifecycle_1.0.4 Rdpack_2.6.4
[91] httpuv_1.6.16 QuickJSR_1.8.1 R6_2.6.1
[94] promises_1.5.0 gridExtra_2.3 codetools_0.2-20
[97] boot_1.3-32 colourpicker_1.3.0 MASS_7.3-58.2
[100] gtools_3.9.5 withr_3.0.2 shinystan_2.6.0
[103] parallel_4.2.3 hms_1.1.4 grid_4.2.3
[106] coda_0.19-4.1 minqa_1.2.8 rmarkdown_2.30
[109] S7_0.2.1 carData_3.0-5 otel_0.2.0
[112] ggforce_0.5.0 shiny_1.11.1 base64enc_0.1-3
[115] dygraphs_1.1.1.6